by: Amadou Soumare
Introduction
The Consumer Price Index (CPI) is one of Canada’s most important economic indicators. Its purpose is to measure the changing cost of a basket of goods and services that are typically purchased by Canadians.
With the publication of the February 2013 CPI on March 27, 2013, significant changes to the CPI took effect. Along with a revised weighting pattern, whereby 2009 expenditure weights were replaced by 2011 weights, this release marked the first time the basket weights were revised at a two-year interval. In addition, the new weights were introduced three months earlier than they were at the time of the previous basket update. These changes, combined with other methodological improvements, have enhanced the quality of the CPI.
The Consumer Price Index
The Consumer Price Index (CPI) is an indicator of the changes in consumer prices experienced by the Canadian population. It compares, through time, the cost of a fixed basket of goods and services purchased by consumers.
The Consumer Price Index is of interest to a wide range of users. It is one of the most widely-known, quoted and utilized statistical series in Canada. Consumers, for example, can compare movements in the CPI to changes in their personal income to monitor and evaluate changes in their financial situation. The CPI is used for economic analysis and research on various issues by business analysts and economists. Private and public pension programs, income tax deductions, and some government social payments are adjusted using the CPI. The index is used as a deflator of various economic aggregates to obtain estimates at constant prices. Finally, the CPI is also a tool for setting and monitoring economic policy. For example, the Bank of Canada uses the CPI and special aggregates of the CPI to monitor its monetary policies.
The Consumer Price Index is a weighted average of the price changes of a basket of goods and services that is based on the expenditure of a target populationNote 1 in a certain reference period. Each good or service is considered as an element in a basket representative of consumer spending, and price movements are assigned a basket share that is proportional to the consumption expenditure for which they account. The weights of goods and services are fixed during the life of a given basket. They play an important role in determining the impact of a given product’s price change on the CPI. For example, Canadians spend a much larger share of their total expenditures on rent than on milk. As a result, a 10% price increase in rental rates will have a greater impact on the All-items CPI than a 10% increase in the price of milk.Note 2
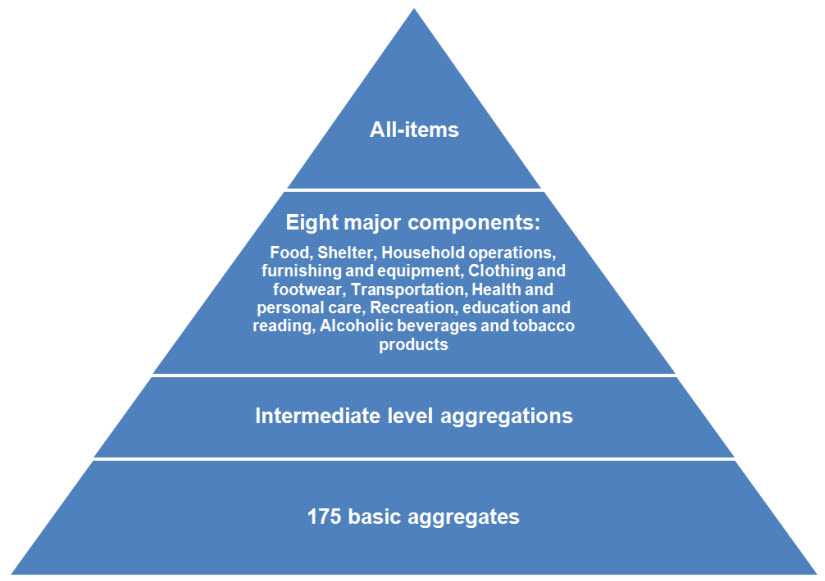
The Consumer Price Index classification of goods and services is organized according to a top-down hierarchical structure (see pyramid). At the top of the structure is the All-items CPI, below which are eight major components. The major components are particularly useful for analytical purposes since they provide a valuable indication about the sources of monthly and annual inflation. At the lowest level of this classification system, there are 175 basic aggregates, which are the building blocks of the CPI.Note 3
Description for Figure
The Consumer Price Index classification is organized according to a top-down hierarchal structure, depicted in a pyramid chart with four levels. At the first level, or the top of the pyramid, is the “All-items Consumer Price Index”. Below at the second level of the pyramid are the eight major components which are:
- Food;
- Shelter;
- Household operations, furnishings and equipment;
- Clothing and footwear;
- Transportation;
- Health and personal care;
- Recreation, education and reading;
- Alcohol beverages and tobacco products.
At the third level of the pyramid there are “Intermediate level aggregations”.
At the fourth and lowest level of the pyramid there are “175 basic classes”.
Consumers modify their purchasing patterns for several reasons, such as changing prices or the arrival of new products on the market. The basket is updated periodically to reflect these changes. The continuity of the CPI series is maintained by chain linking the corresponding indexes obtained from consecutive baskets. This is done separately for each aggregate series, whether it is defined by a product category or by a geographic area. Data users who would like to reconstruct the indexes, or to derive special-purpose indexes consistent with the CPI methodology, should ensure that they include the step of reproducing CPI linking operations.
The time base of an index is the period in which the index is equal to 100. For the Canadian CPI, the time base is usually a calendar year and is expressed as “index year=100.” The current time base is “index 2002=100.”
Importance of Updating the Consumer Price Index Basket Weights
The Canadian CPI is calculated as the change in the price of a specified basket of goods and services. As a Laspeyres-type price index, it is an index in which quantities of the basket are those of the reference period of the weights.
If the fixed basket of goods and services is kept unchanged for an extensive period of time, it will become out of date or irrelevant. This is due, in part, to the fact that consumption patterns have a tendency to evolve in response to shifts in relative prices. Consumer spending patterns are also influenced by factors such as the variation in the level and distribution of household income, demographics (such as an ageing population) and technological changes. New products can also be introduced and existing ones may be modified or become obsolete. As a result, the basket needs to be revised periodically to reflect change in consumers’ spending patterns. For example, the significant increase of the basket share for internet service access from 0.3% in 2001 to 0.8% in 2011 reflects the growing importance of the internet in the daily lives of Canadians.Note 4
In the past, the basket for the Canadian CPI was updated every four to five years using the most current expenditure data from the Survey of Household Spending (SHS). Starting with the 2011 basket update, the weights of the Canadian CPI are now updated every two years. While there are no rules as to how often a CPI basket should be updated, there is a general consensus among CPI experts that more frequent basket updates are preferred to reflect changing consumer spending.
In addition to the review of the expenditure weights, a basket update is also an opportunity to review and update other aspects of the indexes. These may include, but are not limited to:
- Changing the product classification to make it more representative of consumer spending and the products and services available for purchase;
- Reviewing and updating the sample of prices collected;
- Reviewing some product price index estimation methodologies; and
- Updating documentation and dissemination products.
Overview of the 2011 Basket Update
As a fixed-basket Laspeyres-type index, the Canadian CPI is subject to an upward bias brought on by product substitutions made by consumers. This occurs because the index uses a fixed quantity basket from a period in the past and consumers in the intervening months and years change their purchasing behaviour in response to relative price changes of products and services in the marketplace. For example, if the price of chicken increases between basket updates, consumers may opt away from chicken and substitute other meats such as beef. In cases such as these, a fixed-quantity index like the Laspeyres-type price index cannot reflect this expenditure change until the basket weights are updated. This can lead to an overstatement of the importance of changes in the price of chicken in the index and, hence, an upward bias. Typically, the longer a fixed set of basket weights is used, the greater this upward bias.
In 2010, Statistics Canada initiated a number of important changes to the Survey of Household Spending (SHS), the main source of weights for the CPI. Before the redesign, the collection of SHS data was done between January and March following the end of each reference year, with an annual recall period for all spending categories. For example, in the first quarter of 2010, an interviewer visited each household surveyed in the sample and collected data pertaining to the family expenditures in the previous calendar year (2009) for food, shelter, health care, etc. This recall period was determined to be too long for respondents to provide reliable spending estimates for some product groups.
The newly redesigned survey (SHS-R) corrects this weakness by relying on a continuous year-long collection process. Each month, a sample of households is interviewed using recall periods of different lengths for different types of products. Significant, infrequent spending, like the purchase of passenger vehicles, continues to use the 12-month recall period. Other expenditure categories, such as clothing, now use a 3-month recall period. Recurring expenses, for example on rent or utilities, are estimated using the last payment. The SHS-R also introduced a two-week diary for the collection of spending on frequent purchases such as food, gasoline, personal care and alcohol. These changes were implemented in the ten provinces. In the Territories, spending information continues to be collected every two years using the older version of SHS.Note 5
As mentioned earlier, another change with this CPI basket update pertains not only to the frequency of updating the contents and weights of the basket, but also to the time taken to complete this exercise. The frequency and speed of these updates together constitute one of six main criteria used by Statistics Canada to measure data quality. The time between the end of the new reference period and the introduction of the updated CPI basket was shortened. For the 2011 basket update, the new weights took effect 14 months after the end of the new reference period compared to 17 months for the previous basket. This, combined with the effect of more frequent basket updates, translates into a better CPI.
No new product classes were added with this CPI basket update. However, the methodologies of both the prescribed medicines index and purchase of passenger vehicles index were updated and improved. New products were introduced to make the prescribed medicine sample more representative.Note 6 The 2011 basket update also marked the introduction of the new CPI purchase of passenger vehicles classification, basket weights and product sample. Many new car models were added to the CPI sample. Moreover, starting April 2013, the new methodology used to calculate the passenger vehicle parts, maintenance and repair index was introduced, adding new series and replacing less representative products and services.Note 7
Analysis of Basket Weights
The table below shows the 2001 to 2011 CPI basket weights for Canada according to the eight major components and certain product groups. Provincial basket shares for the 2009 and 2011 CPI baskets are provided in Appendix A. The basket share of alcoholic beverages and tobacco products fell the most during this period, from 3.3% in 2001 to 2.8% in 2011. The expenditure share for household operations, furnishings and equipment increased the most, from 11.1% to 12.6%. The basket weight of the health and personal care component also went up, increasing from 4.6% to 5.0%. Over the last decade, shelter costs represented the highest expenditure share of baskets (approximately 26%), followed by transportation expenditures (approximately 20%).
Table 1
Consumer Price Index (CPI) Basket Shares by Major Component and Selected product groups, Canada 2001, 2005, 2009 and 2011 CPI Baskets
Table summary
This table displays the results of Consumer Price Index (CPI) Basket Shares by Major Component and Selected product groups. The information is grouped by Major components, selected product groups (appearing as row headers), Basket weight for (appearing as column headers).
| Major components, selected product groups |
Basket weight for |
|---|
| 2001 |
2005 |
2009 |
2011 |
|---|
| Food |
16.8 |
16.9 |
16.1 |
16.4 |
|---|
| Food purchased from stores |
11.7 |
11.7 |
11.2 |
11.5 |
|---|
| Food purchased from restaurants |
5.1 |
5.1 |
4.8 |
4.9 |
|---|
| Shelter |
26.3 |
25.7 |
27.5 |
25.9 |
|---|
| Rent |
6.1 |
5.3 |
6.0 |
5.8 |
|---|
| Mortgage interest cost |
5.6 |
5.2 |
5.8 |
4.1 |
|---|
| Homeowners’ replacement cost |
3.0 |
3.0 |
4.1 |
4.3 |
|---|
| Property taxes (including special charges) |
3.2 |
3.3 |
3.2 |
3.3 |
|---|
| Water, fuel and electricity |
4.5 |
4.7 |
4.5 |
4.4 |
|---|
| Household operations, furnishings and equipment |
11.1 |
11.4 |
11.8 |
12.6 |
|---|
| Internet access services and subscriptions to online content providers |
0.3 |
0.5 |
0.7 |
0.8 |
|---|
| Clothing and footwear |
6.0 |
5.6 |
5.6 |
6.2 |
|---|
| Transportation |
19.4 |
19.6 |
19.3 |
20.1 |
|---|
| Purchase of passenger vehicles |
7.3 |
6.4 |
6.6 |
6.6 |
|---|
| Gasoline |
3.9 |
4.5 |
4.4 |
4.9 |
|---|
| Health and personal care |
4.6 |
4.8 |
5.0 |
5.0 |
|---|
| Prescribed medicines |
0.6 |
0.6 |
0.6 |
0.9 |
|---|
| Personal care |
2.4 |
2.4 |
2.4 |
2.0 |
|---|
| Recreation, education and reading |
12.5 |
13.0 |
11.8 |
11.3 |
|---|
| Computer equipment, software and supplies |
0.8 |
0.8 |
0.7 |
0.6 |
|---|
| Multipurpose digital devices |
Note ..: not available for a specific reference period |
Note ..: not available for a specific reference period |
0.0 |
0.1 |
|---|
| Home entertainment equipment, parts and services |
1.4 |
1.4 |
1.3 |
0.9 |
|---|
| Alcoholic beverages and tobacco products |
3.3 |
3.1 |
3.0 |
2.8 |
|---|
Changes in basket shares tend to result from evolving social and economic factors. For example, the decreasing importance of cigarettes (down from 1.5% in 2001 to 1.1% in 2011) is reflective of the decline in smoking habits in Canada.Note 8
The important decrease in the share for reading material and other printed material, from 0.7% in 2001 to 0.4% in 2011, was in part attributable to the accessibility to online digital media, which often provided much of the same content at no cost to consumers.
The increasing demand for high-tech goods led to the introduction, in 2009, of a new class for multipurpose digital devices, representing at that time 0.04% of the CPI basket. By 2011, the basket share for this product category had risen to 0.1%, highlighting the continually increasing presence of digital devices such as smartphones and tablet computers in the lives of Canadians.
The ageing Canadian population is likely one of the factors that led to a rise in the importance of medicinal and pharmaceutical products in the CPI basket. The weights for health care goods, for example, grew from 1.0% in 2001 to 1.9% in 2011 as the proportion of the population aged 50 or older reached 34.9%, up from 28.9% in 2001.Note 9
As for the changes between 2009 and 2011, the basket share for food went up from 16.1% to 16.4% as a result of consumers having to pay more for many food products. Food demand is generally less sensitive to price movement than other products.Note 10 This, in part, can explain why food weights didn’t decrease with price increases.
The basket share for shelter was 25.9% in 2011, down from 27.5% in 2009 with all provinces recording a decrease. The basket weight for mortgage interest cost went from 5.8% in 2009 to 4.1% in 2011. This is due in part to the fact that mortgage interest rates decreased significantly during that period. Data from the Canadian Association of Accredited Mortgage Professionals (CAAMP) indicates that in October 2009, households were paying, on average, interest rates of approximately 4.5%, compared to interest rates of approximately 3.9% in the same period in 2011.Note 11,Note 12
An increase was noted in the importance of the household operations, furnishings and equipment component in the 2011 basket. This component reached 12.6% of basket share, up from its previous share of 11.8%. The basket shares for both telephone services and household equipment went up in most provinces.
Clothing and footwear increased to 6.2% of the basket in 2011, up from 5.6% in 2009. In 2011, retail sales of clothing, footwear and accessories reached $36.8 billion, up from $33.8 billion in 2009.Note 13 In the same period, prices for this component decreased 1.6%, suggesting that consumers were taking advantage of low prices to buy more.
The weight for the transportation component went up in most provinces. At the national level, the transportation component reached 20.1% of basket share, up from 19.3% in 2009. Weights for the purchase and leasing of passenger vehicles went down in many provinces.
Between 2009 and 2011, the total share of the health and personal care component remained stable at 5.0%. However, the health care portion showed an increase in most provinces. For Canada as a whole, it reached 2.9% in 2011, up from 2.6% in 2009. This was due, in large part, to spending for medicinal and pharmaceutical products, which accounted for 1.5% of the basket weight in 2011, compared to 1.0% in 2009.
The relative importance of spending for recreation, education and reading declined between 2009 and 2011. Although it represented 11.8% of the basket in 2009, that component’s weight declined to 11.3% in 2011. In most provinces, a decrease in expenditure share for home entertainment, parts and services and those for spectator entertainment was noticed. The basket share of computer equipment and supplies decreased from 0.7% in 2009 to 0.6% in 2011, which was, in part, due to consumers switching from desktop and laptop computers to tablets.
The importance of the alcoholic beverages and tobacco products component in the CPI basket continued to decline in 2011. Alcoholic beverages and tobacco products accounted for 2.8% of expenditures, down from 3.0% in 2009. Alcoholic beverages served in licensed establishments declined significantly in all provinces. However, alcoholic beverages purchased from stores increased in Newfoundland and Labrador, in Prince Edward Island and in Quebec.
Conclusion
The Consumer Price Index (CPI) is Canada’s most well-known measure of consumer inflation. With the 2011 basket update, important changes aimed at improving its quality were introduced. A new practice of updating the basket more frequently and shortening its completion were initiated. Several other methodological improvements were also implemented to significantly enhance the accuracy of the Canadian CPI. These changes, as well as planned enhancements, will ensure the CPI continues to be a reliable indicator of consumer price change.
Notes
- The target population of the CPI consists of all Canadian families and individuals living in urban and rural private households.
- Statistics Canada. Consumer Price Index, February 2013. The Daily, Catalogue 11-001-X (ISSN 1205-9137). Published March 27, 2013.
- Statistics Canada. The May 2011 CPI Basket Update, based on 2009 Expenditures.
- In 2010, 75.5% of Canadians accessed the internet from home at least once a day, up from 63.7% in just five years. For details see: Statistics Canada. Canadian Internet Usage Survey, Survey No. 4432. CANSIM table 358-0155. Note that the target population for the survey has changed from individuals 18 years of age and older in 2005 to individuals 16 years of age and older in 2007.
- The 2011 Survey of Household Spending (SHS) was not conducted in the Territories. Basket weights for the Territories were mostly updated using 2009 price updated expenditures.
- Statistics Canada. Revision of the Prescribed Medicines Index of the Consumer Price Index (CPI), beginning with the September 2012 CPI.
- Statistics Canada. Revision of the Passenger Vehicle Parts, Maintenance and Repair Index of the Consumer Price Index (CPI), beginning with the April 2013 CPI.
- Data from Health Canada show a decline in the overall smoking rate among Canadians aged 15 years and older from 22% in 2001 to 17% in 2011. Health Canada, Canadian Tobacco Use Monitoring Survey (CTUMS), 2012.
- Statistics Canada. Population by Age and Sex for Canada, Survey No. 3604. CANSIM table 051-0001.
- Many studies have found the price elasticity for food demand to be less important than those for other groups of products. For example, research from the United States Department of Agriculture (USDA) used 2005 data to estimate compensated own-price elasticity for nine broad consumption groups in 144 countries. It found the Food, Beverages and Tobacco group to be the less elastic with an own-price elasticity of -0.3, compared to own-price elasticity for other group ranging between -0.6 and -0.8. For details see: United States Department of Agriculture (USDA). International Evidence on Food Consumption Patterns: An update using 2005 International Comparison Program Data. Technical Bulletin No (TB-1929) 59pp, March 2011.
- Canadian Association of Accredited Mortgage Professionals (CAAMP).2009 Annual State of the Residential Mortgage Market in Canada, November 2009.
- Canadian Association of Accredited Mortgage Professionals (CAAMP).2011 Annual State of the Residential Mortgage Market in Canada, November 2011.
- Statistics Canada. Retail Commodity Survey (RCS), Survey No. 2008, CANSIM table 08-0022.
Appendix A
Consumer Price Index (CPI) Basket Shares by Major Component and Selected Product Groups, Provinces, 2009 and 2011 CPI Baskets
Table summary
This table displays the results of Consumer Price Index (CPI) Basket Shares by Major Component and Selected Product Groups. The information is grouped by Major components, selected product groups (appearing as row headers), Newfoundland
and Labrador, Prince Edward Island, Nova Scotia, New Brunswick, Quebec, Ontario, Manitoba, Saskatchewan, Alberta and British Columbia (appearing as column headers).
| Major components, selected product groups |
N.L. |
P.E.I. |
N.S. |
N.B. |
Que. |
Ont. |
Man. |
Sask. |
Alta. |
B.C. |
|---|
| 2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
2009 |
2011 |
|---|
| Food |
16.8 |
16.4 |
16.6 |
16.7 |
16.4 |
16.5 |
16.5 |
16.5 |
19.2 |
19.0 |
14.8 |
15.0 |
16.0 |
16.4 |
14.6 |
15.2 |
15.1 |
15.6 |
15.6 |
16.9 |
|---|
| Meat |
2.7 |
2.9 |
2.0 |
2.4 |
2.0 |
2.1 |
2.3 |
2.1 |
2.5 |
2.5 |
1.8 |
1.9 |
2.0 |
2.1 |
1.7 |
2.1 |
1.8 |
2.0 |
1.7 |
1.9 |
|---|
| Dairy products and eggs |
1.8 |
1.8 |
2.3 |
2.0 |
2.0 |
2.0 |
1.9 |
1.8 |
2.2 |
2.1 |
1.6 |
1.6 |
1.6 |
1.6 |
1.6 |
1.5 |
1.4 |
1.6 |
1.7 |
1.8 |
|---|
| Fruit, fruit preparations and nuts |
1.5 |
1.2 |
1.5 |
1.4 |
1.3 |
1.3 |
1.5 |
1.2 |
1.6 |
1.6 |
1.4 |
1.2 |
1.2 |
1.3 |
1.1 |
1.1 |
1.2 |
1.3 |
1.3 |
1.4 |
|---|
| Vegetables and vegetable preparations |
1.2 |
1.3 |
1.1 |
1.2 |
1.1 |
1.3 |
1.2 |
1.2 |
1.5 |
1.6 |
1.1 |
1.2 |
1.0 |
1.1 |
0.9 |
1.0 |
1.1 |
1.1 |
1.1 |
1.3 |
|---|
| Shelter |
22.0 |
20.6 |
26.0 |
24.0 |
25.5 |
24.5 |
23.0 |
22.0 |
25.6 |
23.6 |
29.1 |
27.3 |
25.1 |
23.3 |
24.8 |
24.2 |
26.3 |
26.1 |
29.8 |
27.6 |
|---|
| Mortgage interest cost |
4.1 |
3.3 |
4.8 |
3.4 |
4.7 |
3.8 |
4.5 |
3.4 |
4.4 |
3.4 |
6.3 |
4.1 |
4.9 |
3.7 |
4.9 |
3.6 |
6.1 |
5.2 |
7.3 |
4.9 |
|---|
| Household operations, furnishings and equipment |
13.1 |
13.6 |
13.3 |
13.3 |
12.8 |
13.9 |
13.3 |
13.6 |
11.1 |
12.0 |
12.2 |
12.9 |
12.4 |
12.6 |
12.2 |
13.1 |
11.8 |
12.2 |
11.2 |
12.4 |
|---|
| Postal and other communications services |
0.2 |
0.1 |
0.1 |
0.2 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.2 |
|---|
| Internet access services and subscriptions to online content providers |
0.8 |
0.8 |
0.9 |
1.0 |
0.8 |
0.9 |
0.8 |
0.8 |
0.7 |
0.8 |
0.7 |
0.8 |
0.8 |
0.8 |
0.7 |
0.7 |
0.7 |
0.7 |
0.6 |
0.7 |
|---|
| Telephone equipment |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.2 |
0.1 |
0.0 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.0 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
0.1 |
|---|
| Clothing and footwear |
5.9 |
7.7 |
5.2 |
6.6 |
5.0 |
5.8 |
5.1 |
6.2 |
5.5 |
6.3 |
5.9 |
6.4 |
5.6 |
6.4 |
5.4 |
5.8 |
5.7 |
6.3 |
5.2 |
5.6 |
|---|
| Transportation |
22.0 |
22.1 |
19.3 |
20.4 |
20.3 |
20.8 |
21.7 |
23.0 |
19.7 |
19.7 |
19.0 |
20.3 |
20.5 |
22.2 |
22.3 |
22.2 |
20.1 |
20.8 |
17.0 |
17.7 |
|---|
| Purchase and leasing of passenger vehicles |
10.0 |
8.8 |
7.5 |
7.1 |
8.7 |
8.5 |
9.6 |
9.9 |
8.8 |
8.3 |
7.0 |
7.0 |
8.4 |
8.4 |
10.4 |
10.3 |
8.7 |
8.4 |
5.7 |
5.8 |
|---|
| Parking fees |
0.0 |
0.1 |
0.1 |
0.3 |
0.1 |
0.2 |
0.1 |
0.2 |
0.1 |
0.4 |
0.2 |
0.4 |
0.2 |
0.3 |
0.1 |
0.2 |
0.2 |
0.3 |
0.1 |
0.4 |
|---|
| Taxi and other local and commuter transportation services |
0.3 |
0.2 |
0.1 |
0.2 |
0.2 |
0.2 |
0.2 |
0.1 |
0.1 |
0.1 |
0.2 |
0.1 |
0.2 |
0.1 |
0.1 |
0.1 |
0.2 |
0.2 |
0.2 |
0.2 |
|---|
| Health and personal care |
5.1 |
4.7 |
5.6 |
5.2 |
4.6 |
5.2 |
5.1 |
5.2 |
5.3 |
5.6 |
4.8 |
4.7 |
5.3 |
4.9 |
5.1 |
5.4 |
4.9 |
4.6 |
4.9 |
5.0 |
|---|
| Health care |
2.7 |
2.6 |
3.0 |
3.5 |
2.3 |
3.2 |
2.8 |
3.1 |
2.8 |
3.3 |
2.4 |
2.7 |
3.0 |
3.2 |
2.7 |
3.2 |
2.6 |
2.8 |
2.8 |
3.1 |
|---|
| Medicinal and pharmaceutical products |
1.3 |
1.5 |
1.6 |
1.9 |
1.1 |
1.7 |
1.2 |
1.7 |
1.2 |
1.5 |
0.9 |
1.3 |
1.5 |
1.8 |
1.3 |
1.8 |
0.9 |
1.4 |
1.1 |
1.6 |
|---|
| Recreation, education and reading |
11.1 |
10.7 |
10.6 |
10.2 |
11.6 |
10.6 |
12.2 |
10.5 |
10.4 |
10.3 |
11.6 |
11.3 |
12.1 |
11.1 |
12.5 |
11.5 |
12.7 |
11.7 |
13.4 |
12.6 |
|---|
| Home entertainment equipment, parts and services |
1.4 |
0.9 |
1.4 |
0.8 |
1.3 |
0.9 |
1.3 |
0.9 |
1.4 |
0.9 |
1.3 |
0.9 |
1.5 |
0.9 |
1.6 |
0.8 |
1.4 |
0.9 |
1.3 |
0.8 |
|---|
| Travel tours |
0.7 |
0.6 |
0.6 |
0.8 |
0.9 |
0.9 |
0.8 |
0.7 |
1.2 |
1.2 |
1.0 |
0.9 |
1.3 |
0.8 |
0.9 |
0.7 |
0.7 |
0.9 |
0.7 |
0.8 |
|---|
| Spectator entertainment (excluding cablevision and satellite services) |
0.3 |
0.2 |
0.5 |
0.3 |
0.4 |
0.3 |
0.4 |
0.2 |
0.4 |
0.3 |
0.5 |
0.3 |
0.6 |
0.5 |
0.7 |
0.3 |
0.6 |
0.3 |
0.4 |
0.4 |
|---|
| Alcoholic beverages and tobacco products |
3.9 |
4.2 |
3.5 |
3.6 |
3.8 |
2.7 |
3.2 |
3.0 |
3.3 |
3.7 |
2.6 |
2.2 |
3.0 |
3.2 |
3.1 |
2.6 |
3.3 |
2.8 |
3.0 |
2.4 |
|---|
| Alcoholic beverages served in licensed establishments |
0.4 |
0.2 |
0.5 |
0.3 |
0.5 |
0.3 |
0.4 |
0.3 |
0.5 |
0.4 |
0.5 |
0.4 |
0.5 |
0.4 |
0.6 |
0.4 |
0.6 |
0.5 |
0.7 |
0.4 |
|---|
| Alcoholic beverages purchased from stores |
1.4 |
1.6 |
1.1 |
1.3 |
1.3 |
1.0 |
1.2 |
1.1 |
1.4 |
2.0 |
1.2 |
0.9 |
1.0 |
1.0 |
1.1 |
0.7 |
1.1 |
1.0 |
1.3 |
1.1 |
|---|
